This Article
Expression of Cucumber Mosaic Virus Coat Protein and Its Assembly into Virus Like Particles
Abstract
Background: Cucumber mosaic virus (CMV) has isometric particles with a diameter of about 28–29 nm. A useful method to recognize regions from mutagenesis in the coat protein (CP) gene are important for particle assembly and the expression of viral in CP expression systems like bacteria such as E.coli or yeast as well as their assembly into virus-like particles (VLP).
Objectives: In this paper, we report the expression and assembly of VLP in cells of E. coli expressing the CMVCP gene.
Materials and Methods: In this study, the CMV-CP gene was released from a previously prepared cloning vector. Then, the CMV-CP was ligated into the expression vector. Sequencing was done by Marcrogen, Inc. (South Korea). A recombinant plasmid was transferred to E.coli isolate Rosetta. After inducing by isopropyl thiogalactosides, the molecular weight of the expressed protein was determined by SDS-PAGE. The extraction of proteins was done by NATURE method to see the possible presence of CMV-like particles.
Results: CMV-CP was detected by Western blotting by a CMV specific polyclonal antibody and conjugate. The protein extracted from the CP producing clone was studied under a JEOL 100-CXII transmission electron microscope with 100000× magnification at an acceleration voltage of 100 kV.
Conclusions: The results showed that the CP gene was expressed in the prokaryotic system successfully and was assembled into the CMV-like particle.
Keywords: Cucumovirus; Gene Expression; Recombinant Proteins; Virus Assembly; Electron Microscopy
1. Background
Cucumber mosaic virus (CMV) belongs to the genus Cucumovirus in the family Bromoviridae. It is plant virus present worldwide with an extensive host range that infects about 1200 species in almost 85 families of plants including cereals, fruits, vegetables, and ornamentals (1). CMV is a multi-component virus consisting of three single-stranded genomic RNAs and two subgenomic RNAs known as sgRNA4 and sgRNA4A (2–4). The CMV CP is translated from a subgenomic (sg) RNA 4 corresponding to the 3´ proximity of the viral RNA 3 (5). Its particles are isometric with a diameter of about 28–29 nm (4, 6) and the virions consist of 180 identical copies of the virus-encoded coat protein (CP) (5). Proteins encoded by RNAs 1 and 2 are needed for viral genome replication while RNA3 encode the movement protein (MP) and CP, both of which are involved in cell-to-cell and long-distance movement of the virus (4). CMV is transmitted mechanically by sap and aphids in a non-persistent, stylet-borne manner (7).
A useful way to recognize regions in the CP gene that are important for particle assembly from mutagenesis is the expression of viral CPs in the expression systems like bacteria or yeast as well as their assembly into virus-like particles (VLP). Assembly of VLP from heterologously expressed CPs has been studied in detail primarily for the Johnsongrass mosaic virus, Potato virus Y (8–10), Alfalfa mosaic virus (11), and CMV (12). By the molecular assembly of the capsid and production of recombinant antibodies, can be obtained by assembly of VLP in plants or bacteria (13, 14).
Cloning and expression of viral genes has been an important strategy for obtaining large scale recombinant proteins that may be used in cases such as antibody production (15–17) given that diagnosis and detection of plant viruses in a large number of samples is done with serological methods due to their robustness and low cost (18, 19). The cap protein is highly immunogenic and reacts strongly with the antibody. Therefore, it is a good candidate antigen for production recombinant antibody against CMV and for the development of serologic tests. Expression of CPs in microbial systems and their assembly into virus-like particles facilitate large-scale production of CP in vitro, which represents a pure antigen free from contaminating plant proteins that can be used to produce polyclonal antiserum.
2. Objectives
In this paper, we report the expression and assembly of VLP in cells of E. coli expressing the CMVCP gene.
3. Materials and Methods
3.1. Cloning, Sequencing and Expression
The CMVCP gene (from strain B13) was cloned into pTZ57R/T vector and transformed into E. coli strain DH5α, then it was removed from pTZ57CMVCP by digestion with BamH I and Sac I restriction enzymes. The CMV-CP was then ligated into the expression vector pET21a, previously digested with the same enzymes. The recombinant plasmid pET21aCMVCP was transformed into E. coli strain Rosetta with a heat shock procedure. Then, the sequencing was done by Macrogen Inc. (Seoul, South Korea) to confirm the correct insertion of the CP gene in the vector.
A single transformant colony was cultured overnight in 5 mL of LB (1% peptone, 0.5% yeast extract, 1% NaCl) with ampicillin (final concentration of 50 μg/mL), then 1 mL of an overnight culture was added to 50 mL fresh LB ampicillin plus. When the culture had reached an OD (optical density) of 0.6, induced by adding 1 mM isopropyl thiogalactosides (IPTG) and shake-incubated at 37°C. Induction periods of 3–5 hours were applied. Then, cells were harvested by centrifugation (5000 rpm, 10 minutes, 4°C) and stored at -80°C until used.
3.2. SDS-PAGE and Western Blotting
The extracted proteins from the harvested cells (by 160 μL of Laemmli buffer: 125 mM Tris/HCl, pH 6.8, 2% sodium dodecyl sulphate, 9% glycerol, 0.7 M 2-mercaptoethanol, 0.002% bromophenol blue, and boiled for 2–5 minutes) were separated by 12% sodium dodecyl sulphate polyacrylamide gel electrophoresis (SDS-PAGE) for 4 hours at 120 V (20) and visualized by 0.2% Coomassie Blue R staining solution. Then, the protein bands were electro blotted onto a HybondTM-N nitrocellulose membrane (Amersham, UK) using a wet transfer system (Akhtarian, Tabriz) at 100 mA for 1 hour for Western blot analysis. For blocking, we used of 5% (w/v) BSA powder in the buffer (8 mM Na2HPO4, 1.5 mM KH2PO4, 2.7 mM KCl, 137 mM NaCl, pH 7.3) for 1 hour. After washing twice with PBST (phosphate buffered saline- tween 20) membranes were incubated in 1:1000 dilution of polyclonal anti-CMV antibodies (Agdia, USA) for 2 hours at room temperature, followed by washing as above and incubating in alkaline phosphatase-conjugated anti-rabbit IgG for 2 hours at room temperature. After washing three times, the CMV CP specific band was visualized by incubating the membrane in the substrate NBT/BCIP (nitroblue tetrazolium chloride 5-bromo-4-chloro-3-indolyl-phosphate, toluidine salt, Roche) solution.
3.3. Transmission Electron Microscopy
The extraction protocol that we used to extract soluble protein from bacteria was from Folwarczna et al. (21) and was optimized for our usage. For Transmission electron microscope (TEM) studies, the bacterial cells that were harvested by centrifugation, resuspended in 1 mL of TE, and incubated at 20°C for 5–10 minutes with lysozyme (1 mg/mL). Then for the lysing of bacteria, osmotic shock was applied by solubilizing in 1 vol of 40% sucrose (wt/vol), followed by adding 4 vol of TE buffer. DNase and RNase was added in final concentration 5 Mg/mL and 10 mg/mL, respectively. Finally, the cell debris was removed by centrifuge in 12000 rpm and 4°C for 20 minutes, and then the supernatant were analyzed by TEM.
A drop of the CMVCP recombinant protein extracted from the bacteria as described above was adsorbed onto a carbon shadowed formvar coated grid for 2–3 hours at room temperature and diluted in 0.06 M phosphate buffer, pH 6.5.Then, the grids were dried gently using filter paper. The grid was negatively stained by 2% uranyl acetate for 30 seconds. It was studied under a JEOL 100-CXII transmission electron microscope with 100000× magnification at an acceleration voltage of 100 kV.
4. Results
4.1. Preparation of Construct and Expression of Coat Protein in E. coli
The sequencing confirmed correct insertion after ligation of the CP cDNA into pET2la. The insertion was in frame and no stop codon was created; however, additional nucleotides were inserted into the CP cDNA because of the restriction sites used in the cloning. Analysis by SDS-PAGE of the total protein from bacterial cells carrying pET21aCMVCP and induced by IPTG in the range of 0.5–2 mM (final concentration) for 4 hours showed a higher level expression of the expected protein of approximately 25 kDa in size corresponding to the CMV CP (12) (Figure 1A). Four independent transformed (clones) were examined for the expression among which two clones with a desired level of the expression were selected for further analysis.
Western blotting with a rabbit anti-CMV polyclonal antibody confirmed the identity of the recombinant protein as CMVCP. Accordingly, a protein band with a molecular weight of 26 kDa (the expected size (based on comparison with the protein size marker)) was observed in the Western blot (Figure 1B).
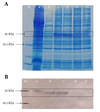 |
SDS-PAGE Analysis and Western Blotting Results
|
4.2. In Vitro Assembly of CMVCP
Plant virus CP genes expression in the bacteria can assemble into virus like particles. Microscope observation of the CMVCP protein extracted from induced bacteria was studied by TEM in a magnification of 100000×. The results of this study show that formation of CMV-like particles had been carried in the E. coli bacterial system.
The results of the assembled recombinant CMVCP into CMV-like particles suggests that CMVCP recombinant protein expressed in the E. coli system can assemble into VLPs (Figure 2).
|
|
Transmission Electron Microscope of Extracted Protein
|
5. Discussion
E. coli strain Rosetta belongs to so-called “B strains”, which are deficient in the lon protease and lack the ompT outer membrane protease. The strains having these proteases can degrade proteins during purification (22). Therefore, strain Rosetta has been the cell line of choice for expression of genes of interest (13, 23). Also, this strain (Rosetta) is optimal when the target gene contains rare codons or the expressed protein has disulfide bonds (24). In our study, we tested IPTG concentrations ranging from 0.5–2 mM and found that changes in this range do not affect expression levels. It has been reported that the duration for expression of the gene of interest at a particular IPTG concentration may vary according to the gene of interest or other factors. Here, we used induction periods of 3–5 hours and found that an induction period of 4 hours gave the best result (data not shown).
This research results indicated that the VLPs of the CMVCP protein were successfully expressed in E. coli and the CMVCP protein was able to self-assemble into VLPs in vitro. TEM studies showed that shape and size were similar to CMV isolated from the cucurbita leaf infected with CMV. The size of the CMV particles was determined as between 28 nm (4, 6) and 30 nm (25). Given these results, the particles seen by TEM studies are probably CMV like particles.
Accordingly, it has been demonstrated that the coding sequence of the CMV CP open reading frame (ORF) placed under the subgenomic promoter of the potato virus X (PVX) is translated into the CP subunits, which are capable of assembling into CMV virus-like particles (VLPs) in the absence of a full-length RNA 3 molecule and CMV replication machinery associated with RNA 1 and RNA 2.
The self-assembled empty VLPs have the potential to be used as inner constrained environments for the preparation of nano-structured materials that may be promising as potential drug carriers (26). We showed that CMV CP ORF placed under the control of T7 promoter in the bacterial expression vector pET-21a(+) can be expressed to form virus like particle in the bacterium. Previous studies reported production of VLPs for carrying nano-materials, including Cowpea chlorotic mottle virus, Tomato bushy stunt virus, and Turnip crinkle virus (27–29).
VLPs mimic the structure of authentic virus particles and present viral antigens in a more authentic conformation and biological function. Therefore, they are easily recognized by the immune system and able to stimulate B-cell and T-cell immune responses. In particular, many VLPs are non-infectious, because, although they completely lack the viral DNA or RNA genome, they are safer than attenuated or chemically inactivated live viruses. This striking feature of VLPs will likely contribute to the effectiveness of its use for production recombinant antibody.
This study demonstrated the achievability of expression of the CP from an Iran isolate of the virus (CMV-B13). An antibody prepared against an indigenous isolate may perform efficiently to screen for a virus in the same locality. This is because viruses undergo steady variation and the virus CP gene is not exceptional. Accordingly, commercial antibodies, which are prepared against a foreign isolate of a given virus, may not be efficient enough to detect the virus in another locality.
Footnotes
References
- 1. Roossinck MJ. Cucumber mosaic virus, a model for RNA virus evolution. Mol Plant Pathol. 2001;2(2):59-63. [PubMed]
- 2. Palukaitis P, Garcia-Arenal F. Cucumoviruses. Adv Virus Res. 2003;62:241-323. [PubMed]
- 3. International Committee On Taxonomy Of Viruses , Van Regenmortel HV, Fauquet CM, International Union of Microbiological Societies, Virology Division , Bishop DHL. International Committee On Taxonomy Of Viruses. Virus Taxonomy: Classification and Nomenclature of Viruses : Seventh Report of the International Committee on Taxonomy of Viruses. Academic Press; 2000. ISBN 9780123702005.
- 4. Palukaitis P, Roossinck MJ, Dietzgen RG, Francki RI. Cucumber mosaic virus. Adv Virus Res. 1992;41:281-348. [PubMed]
- 5. Smith TJ, Chase E, Schmidt T, Perry KL. The structure of cucumber mosaic virus and comparison to cowpea chlorotic mottle virus. J Virol. 2000;74(16):7578-86. [PubMed]
- 6. Madhubala R, Bhadramurthy V, Bhat AI, Hareesh PS, Retheesh ST, Bhai RS. Occurrence of Cucumber mosaic virus on vanilla (Vanilla planifolia Andrews) in India. J Biosci. 2005;30(3):339-50. [PubMed]
- 7. Ng JC, Liu S, Perry KL. Cucumber mosaic virus mutants with altered physical properties and defective in aphid vector transmission. Virology. 2000;276(2):395-403. [PubMed]
- 8. Jagadish MN, Hamilton RC, Fernandez CS, Schoofs P, Davern KM, Kalnins H, et al. High level production of hybrid potyvirus-like particles carrying repetitive copies of foreign antigens in Escherichia coli. Biotechnology (N Y). 1993;11(10):1166-70. [PubMed]
- 9. Jagadish MN, Ward CW, Gough KH, Tulloch PA, Whittaker LA, Shukla DD. Expression of potyvirus coat protein in Escherichia coli and yeast and its assembly into virus-like particles. J Gen Virol. 1991;72 ( Pt 7):1543-50. [PubMed]
- 10. Stram Y, Yadin H, Chai D, Molad T. Identification of foot and mouth disease virus (FMDV) serotypes using the polymerase chain reaction. Israel J Veterinary Med. 1993;48-57.
- 11. Yusibov V, Kumar A, North A, Johnson JE, Loesch-Fries LS. Purification, characterization, assembly and crystallization of assembled alfalfa mosaic virus coat protein expressed in Escherichia coli. J Gen Virol. 1996;77 ( Pt 4):567-73. [PubMed]
- 12. Natilla A, Hammond RW, Nemchinov LG. Epitope presentation system based on cucumber mosaic virus coat protein expressed from a potato virus X-based vector. Arch Virol. 2006;151(7):1373-86. [PubMed]
- 13. Bragard C, Duncan GH, Wesley SV, Naidu RA, Mayo MA. Virus-like particles assemble in plants and bacteria expressing the coat protein gene of Indian peanut clump virus. J Gen Virol. 2000;81(Pt 1):267-72. [PubMed]
- 14. Hwang DJ, Roberts IM, Wilson TM. Expression of tobacco mosaic virus coat protein and assembly of pseudovirus particles in Escherichia coli. Proc Natl Acad Sci U S A. 1994;91(19):9067-71. [PubMed]
- 15. Barbieri MR, Carvalho MGD, Zambolim EM, Zerbini FM. Expression in Escherichia coli of the capsid protein of Watermelon mosaic virus and production of specific antiserum. Fitopatologia Brasileira. 2004;29(2):215-9.
- 16. Nickel O, Targon MLPN, Fajardo TVM, Machado MA, Trivilin AP. Polyclonal antibodies to the coat protein of Apple stem grooving virus expressed in Escherichia coli: production and use in immunodiagnosis. Fitopatologia Brasileira. 2004;29(5):558-62.
- 17. Targon M, Nikolaeva O, Manjunath KL, Lee RF, Mueller GW, Machado MA. Coat protein gene of a Brazilian isolate of the Citrus tristeza virus: cloning, expression in E. coli and production of polyclonal antiserum. Fitopatologia Brasileira. 1997;22:99-102.
- 18. Ling KS, Zhu HY, Jiang ZY, Gonsalves D. Effective Application of DAS-ELISA for Detection of Grapevine Leafroll Associated Closterovirus-3 Using a Polyclonal Antiserum Developed from Recombinant Coat Protein. Eur J Plant Pathol. 2000;106(4):301-9. [DOI]
- 19. Zimmermann D, Bass P, Legin R, Walter B. Characterization and Serological Detection of Four Closterovirus-like Particles Associated with Leafroll Disease on Grapevine. J Phytopathology. 1990;130(3):205-18. [DOI]
- 20. Laemmli UK. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 1970;227(5259):680-5. [PubMed]
- 21. Folwarczna J, Plchova H, Moravec T, Hoffmeisterova H, Dedic P, Cerovska N. Production of polyclonal antibodies to a recombinant coat protein of potato virus Y. Folia Microbiol (Praha). 2008;53(5):438-42. [PubMed]
- 22. Grodberg J, Dunn JJ. ompT encodes the Escherichia coli outer membrane protease that cleaves T7 RNA polymerase during purification. J Bacteriol. 1988;170(3):1245-53. [PubMed]
- 23. Jacob T, Usha R. Expression of Cardamom mosaic virus coat protein in Escherichia coli and its assembly into filamentous aggregates. Virus Res. 2002;86(1-2):133-41. [PubMed]
- 24. Novagen pET System Manual. 11th ed. Gibbstown, N. J.: Novagen (EMD Biosciences); 2006/2007.
- 25. Kim JH, Choi GS, Choi JK. Characterization of Cucumber mosaic virus Subgroup 2 Isolated from Parkika(Capsicum annuum var.grossum)in Korea. Plant Pathol J. 2002;18(1):6-11.
- 26. Douglas T, Young M. Virus Particles as Templates for Materials Synthesis. Adv Mater. 1999;11(8):679-81. [DOI]
- 27. Tama F, Brooks CL, 3rd.. The mechanism and pathway of pH induced swelling in cowpea chlorotic mottle virus. J Mol Biol. 2002;318(3):733-47. [PubMed]
- 28. Harrison BD, Robinson DJ. Genome reconstitution and nucleic acid hybridization as methods of identifying particle-deficient isolates of tobacco rattle virus in potato plants with stem-mottle disease. J Virol Methods. 1982;5(5-6):255-65. [PubMed]
- 29. Sorger PK, Stockley PG, Harrison SC. Structure and assembly of turnip crinkle virus. II. Mechanism of reassembly in vitro. J Mol Biol. 1986;191(4):639-58. [PubMed]

