This Article
Except where otherwise noted, this work is licensed under Creative Commons Attribution-NonCommercial 4.0 International License.
Evaluation of AmpC Gene Expression in Carbapenem Resistant Pseudomonas aeruginosa Strains Isolated from Severely Burned Patients With Secondary Infection in Hospitals
Received: Nov 9, 2014;
Accepted: Nov 20, 2014;
epub: Nov 25, 2014;
ppub: Nov 2014
Abstract
Background: Pseudomonas aeruginosa is an opportunistic pathogen mostly affecting hospitalized and immunodeficient patients. Considering the extent of attention paid to the issue of antibiotic resistance and the origin of such resistance and the growing number of treatment failures in patients with burn injury, it seems that investigating the degree of ampC gene expression could be beneficial for the treatment of patients with severe burn injury.
Objectives: The present study focused on identifying different antibiotic patterns, detecting carbapenem resistant Pseudomonas aeruginosa strains isolated from severely burned patients and investigating AmpC gene expression as one of the important mechanisms associated with drug resistance.
Materials and Methods: A total of 189 clinical isolates of carbapenem resistant Pseudomonas aeruginosa, isolated from patients with severe burn injuries, were identified by bacteriological methods followed by determination of their antibiotic resistance patterns by the standard protocol of Kirby–Bauer. The expression of ampC gene was determined by quantitative real-time PCR.
Results: Based on our findings, 94.2% of the isolates were resistant to imipenem, 99.5% were resistant to meropenem, and all were resistant to ertapenem. The level of ampC gene expression in carbapenem resistant Pseudomonas aeruginosa isolates, compared to standard carbapenem sensitive strains, was significantly increased.
Conclusions: Our data showed that the over-expression of ampC gene is an important mechanism of resistance in carbapenem resistant Pseudomonas aeruginosa strains isolated from severely burned patients with secondary infections.
Keywords: Pseudomonas aeruginosa; AmpC beta-lactamases; Drug Resistance
1. Background
Pseudomonas aeruginosa is an opportunistic, aerobic gram-negative pathogen that possesses lipopolysaccharides, a single polar flagellum and pili. Flagellum, the motility organ of bacteria, confers bacteria the ability to change direction and is important for attachment of bacteria to cell membranes thus, plays a vital role, as a virulence factor, in hospitalized immunodeficient patients (1). The ability of bacteria in developing β-lactamases leads to the emergence of the resistance phenomenon in many species of microorganisms, in particular those responsible for nosocomial infections, making treatment strategies for such infections extremely complicated (2). The spectrum of β-lactamase activity of some bacteria may expand, leading to the appearance of extended spectrum β-lactamase (ESBL)-producing organisms. Bacterial β-lactamases are divided into four major groups (A-D) according to the Ambler classification scheme. Group A hydrolyze penicillin and cephalosporins; group B are metallo-β-lactamases (MBLs) requiring Zn2+ ion(s) for activity; group C include enzymes such as AmpC-type β-lactamases (3) that degrade cephamycins and cephalosporins; and group D are β-lactamases with high hydrolysis rate. AmpC-induced resistance is of serious concern in Pseudomonas aeruginosa (4). Resistance to carbapenems in Pseudomonas aeruginosa may be due to the production of AmpC-type β-lactamases (5). Also, AmpC-type β-lactamases cause bacteria to become resistant to aztreonam and inhibit class A enzymes such as clavulanic acid. The most relevant mechanism for expanding resistance to antipseudomonal penicillins and cephalosporins is the occurrence of mutations that produce chromosomally mediated AmpC β-lactamases with the potential to hydrolyze cephalosporins. These mutations occur during the course of antibacterial therapy, leading to treatment failure (6). Despite numerous advances in treating patients with severe burn injuries, these injuries are still one of the most important causes of mortality throughout the world with burn wound infections and secondary sepsis as the most important complication following major burn injuries (7). Most deaths (75%) in patients with severe burns of greater than 40% of the total body surface are currently associated with blood infections and burn wound infections or other infection complications and respiratory damages. Also, burn injury damages the first line of the host defense barriers and suppresses the body defense system, facilitating the entry of opportunistic and pathogenic organisms such as Pseudomonas aeruginosa and Staphylococcus aureus into the body.
Upon injury, the initial colonization occurs with gram-positive and gram-negative bacteria from the patient’s endogenous skin flora and later by microorganisms transmitted from the hospital environment (8). Vancomycin-resistant enterococci (VRE), methicillin-resistant Staphylococcus aureus (MRSA), and multidrug-resistant gram-negative bacteria (MDRGN) which contain several types of β-lactamases including extended spectrum β-lactamases (ESBLs), AmpC β-lactamases and metallo-β-lactamases are among the most important pathogenic bacteria affecting hospitalized patients (9).
Considering the intrinsic and acquired properties of Pseudomonas aeruginosa, compared to enterobacteriaceae, these organisms are resistant to many antibacterial agents. Antibacterial treatment against these organisms is performed by administrating synthetic penicillins, third-generation cephalosporins, carbapenems, monobactams, aminoglycosides, fluoroquinolones and polymyxins; commonly a combination of an aminoglycoside plus a β-lactam drug, third-generation cephalosporin, monobactams or carbapenem is preferred. However, combination therapy is a matter of controversy and has been emphasized that mono-therapy could be as effective as combined therapy if an appropriate antibacterial agent is administered at the correct dose and the right time (10). Carbapenems are the drug of choice when fighting ESBLs-producing Pseudomonas aeruginosa because of their high-level of stability against the hydrolytic activity of ESBLs, small size, and their double-ring structure which eases their passage through the bacterial outer membrane (11). However, the rate of treatment failure and mortality following the administration of carbapenems is still high while, the development of resistance during therapy against Pseudomonas aeruginosa infections occurs in many cases, depending on the drug dosage, type of drug used, and the location of infection. Among carbapenems, there is greater possibility for developing resistance against imipenem during the course of treatment (12). The emergence of resistance against imipenem in Pseudomonas aeruginosa seems to be in coordination with the loss of oprD and AmpC β-lactamase activity (13). Clinicians recommend that the initial treatment should be started with a combination of an antipseudomonal β-lactam drug plus an aminoglycoside (12).
2. Objectives
The present study focused on identifying different antibiotic patterns, detecting carbapenem resistant Pseudomonas aeruginosa strains isolated from severely burned patients, and investigating AmpC gene expression as one of the important mechanisms associated with drug resistance.
3. Materials and Methods
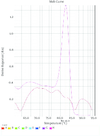
A total 189 clinical isolates of carbapenem-resistant P. aeruginosa were collected from different burn units in Tehran (Iran) during 2011 and 2012. Clinical isolates were confirmed by standard bacteriological methods performed at the medical microbiology laboratory of Qazvin University of Medical Sciences. Briefly, the organisms included in the study were gram-negative obligate aerobic bacteria with a positive oxidase test, inability to ferment glucose or lactose under anaerobic conditions in oxidation/fermentation medium (OF), and ability to grow on different culture media such as blood agar, MacConkey agar, eosin methylene blue agar, and cetrimide agar. The antibiotic-resistance profile of isolated organisms was evaluated by the Kirby Bauer disk diffusion assay according to the Clinical and Laboratory Standards Institute (CLSI) guidelines. The antibiotics used were imipenem (10 µg), meropenem (10 µg), ertapenem (10 µg); all obtained from the MAST Company (England). A standard strain of P. aeruginosa (ATCC 27853) was used as the control organism. To investigate the level of ampC β-lactamase gene expression, a quantitative real-time polymerase chain reaction (qRT-PCR) assay was employed. For RNA extraction and performance of RT-PCR, P. aeruginosa isolates were cultured in 1.5 mL lysogeny broth (LB) and left in a shaking incubator (180 rpm) at 37 оC for 18-24 hours. Later, following overnight incubation, the cultures were centrifuged (8000 rpm) at 4 оC for seven minutes. The sediment obtained was used for extraction of total RNA using a commercial kit (RNeasy kit, Qiagen, Germany), according to the manufacturer’s instructions. The elimination of genomic DNA was achieved using a commercial RNase-free DNase kit (DNase |, RNase-free, Fermentas, Thermo Scientific, USA) according to the manufacturer’s instructions. To prepare cDNA, 5 µL of extracted RNA was added to 15 µL distilled, deionized water and converted to cDNA in a total volume of 20 µL using a commercial kit (cDNA kit, BioNeer Pacific, Australia) according to the manufacturer’s instructions. The quality and quantity of extracted RNA was assessed by electrophoresis on agarose gel. Real-time PCR was performed using the Applied Biosystems 7500 real-time PCR (Fort Collins, Colorado, USA) and a commercial kit (SYBR® Green2X Master Mix; Applied Biosystems, USA). Briefly, using a standard 96-well microplate (Applied Biosystems; USA), a volume of 0.5 µL of each primer, 5 µL cDNA, 4 µL distilled deionized water, and 10 µL of Master Mix were added to each well. All PCR reactions were performed in a 20 µL volume for 45 cycles. After PCR performance, a melting curve was drawn to evaluate the specificity of the PCR reaction (Figures 1 and 2) followed by electrophoresis of PCR products on 2% agarose gel (Figure 3). Specific primers of ampC (Table 1) were used to evaluate changes of ampC gene expression in carbapenem-resistant P. aeruginosa isolates. Housekeeping gene rpoD was used as an internal standard for normalizing the level of transcription of target genes and the standard P. aeruginosa strain (ATCC 27853) for comparative evaluation of the gene mentioned above.
|
|
ampC Gene Linear Amplification Plot
|
|
|
Melting Curve Plot for the Isolates With ampC gene
|
|
|
PCR Product
|
|
Table 1. Primers Used to Evaluate the ampC β-lactamase Gene Expression
|
3.1. Method Used to Evaluate Gene Expression
Once the PCR reaction was over, the threshold line was determined and the data were analyzed by descriptive statistics using the Microsoft Excel software. To determine the output and correlation coefficient of the target gene (ampC) and the housekeeping gene (rpoD) from extracted RNA, serial dilutions were prepared and following performance of RT-PCR, a standard curve was drawn for each segment of the gene. Later, the CT of samples was used to calculate the increase or decrease in gene expression using the 2-ΔΔCT method.
4. Results
The findings of the drug susceptibility assay using the Kirby-Bauer disk diffusion technique on 189 carbapenem-resistant P. aeruginosa, isolated from burn wounds, revealed 94.2%, 99.5%, and 100% resistance rate towards imipenem, meropenem and ertapenem, respectively. The results of this study indicate that the ampC gene expression was increased in all studied isolates compared to the rpoD housekeeping gene. Of the 189 clinical isolates of Pseudomonas aeruginosa, 94.2%, 99.5%, and 100% were resistant to imipenem, meropenem and ertapenem, respectively.
5. Discussion
Beta-lactam drugs compose more than 50% of all antibiotics used in clinics (14) yet the therapeutic benefits of this group of drugs is decreasing because of the growing prevalence of bacterial resistance mechanisms (15). The eminent resistance mechanism against the β-lactam drugs in Pseudomonas aeruginosa and many other non-fermenting gram-negative bacteria, like many members of the family Enterobacteriaceae, is the production of AmpC β-lactamases (16, 17). The AmpC β-lactamases are responsible for inactivating a broad spectrum of diverse β-lactam drugs such as penicillins, cephalosporins and monobactams (18). The most relevant mechanism for expanding the resistance phenomenon to antipseudomonal penicillins is the occurrence of mutations that lead to hyperproduction of the chromosomal cephalosporinase AmpC. During the course of treatment with β-lactam agents, resistant mutations with constitutive AmpC over-production are commonly selected, resulting in treatment failure (6). The development of resistance against imipenem in Pseudomonas aeruginosa seems to be the interplay between AmpC β-lactamase overproduction and the loss of OprD (13). In the current study, the CT values obtained for ampC and rpoD genes in carbapenem-sensitive Pseudomonas aeruginosa isolates were 31.6422 ± 3.8728 and 32.011 ± 6.1982, respectively, with ΔCT of 0.3688. Likewise, the CT values calculated for ampC and rpoD genes in carbapenem-resistant Pseudomonas aeruginosa strains isolated from patients were 33.9869 ± 3.8728 and 24.3809 ± 5.5262, respectively, with a value of -9.9768 for ΔCT. Considering the formula 2-ΔΔCT, it is clear that the level of ampC gene expression in clinical isolates of carbapenem-resistant Pseudomonas aeruginosa was 1300 times greater than that found for carbapenem-sensitive Pseudomonas aeruginosa used as the standard strain. Quale et al. in 2006 studied 33 isolates of Pseudomonas aeruginosa with different susceptibility patterns against carbapenem and tested these organisms for the expression of ampC, oprD, and efflux genes using RT-PCR. Based on the RT-PCR findings, these clinical strains were divided into three groups among those, 13 isolates from group one showed AmpC overproduction (19). Also, Cabot et al. in 2011 evaluated the prevalence and effect of AmpC and efflux pump overexpression in 190 clinical strains of Pseudomonas aeruginosa isolated from blood samples using real-time PCR and showed AmpC overproduction in 24.2% of isolates (20). In another study by Xavier et al. in 2010, the level of expression for efflux pump, oprD, and ampC genes in Pseudomonas aeruginosa strains was evaluated in which 52.5% of isolates were resistant to carbapenems while 11.9% showed no Ampc overproduction (21). Finally, Tomas et al. in 2010 examined 25 clinical strains of Pseudomonas aeruginosa, isolated from patients with cystic fibrosis, for the expression of ampC, oprD, mexA, mexC, mexE, and mexX genes and demonstrated an overexpression for the mexA and ampC genes yet a reduced level of expression for the oprD gene, a finding correlated with resistance against β-lactams (22).
Considering the extent of attention paid to the issue of antibiotic resistance and the origin of such resistance and as there is a growing number of treatment failures in patients with burn injury, it seems that investigating the degree of ampC gene expression could be beneficial for the treatment of patients with severe burn injury. The present study focused on identifying different antibiotic patterns, detecting carbapenem resistant Pseudomonas aeruginosa strains isolated from severely burned patients, and investigating the AmpC gene expression as one of the important mechanisms associated with drug resistance. This is the first report from Iran concerning one of the resistance mechanisms against carbapenem, i.e. AmpC β-lactamase expression in Pseudomonas aeruginosa. On a nationwide scale, the findings of this study are of crucial importance and could be implemented in broadening a new strategy for antipseudomonal treatment.
Changes in the strategy of antibiotic administration and establishing appropriate infection control measures in hospital wards, in particular those designed for long-term hospitalization such as burn units, are important factors that could, to some extent, prevent the dissemination of carbapenem-resistant organisms.
Acknowledgments
The authors of the present study gratefully thank and acknowledge their colleagues working at the Department of Microbiology, the Basic Sciences Research Laboratory, and the Reference Laboratory of Qazvin University of Medical Sciences, Qazvin, Iran for their sincere assistance with this research project.
Footnotes
References
- 1.
Driscoll JA, Brody SL, Kollef MH. The epidemiology, pathogenesis and treatment of Pseudomonas aeruginosa infections. Drugs. 2007;67(3):351-68. [PubMed] - 2.
Paterson DL. Resistance in gram-negative bacteria: enterobacteriaceae. Am J Med. 2006;119(6 Suppl 1):S20-8. discussion S62-70 [PubMed] - 3.
Jacoby GA, Munoz-Price LS. The new beta-lactamases. N Engl J Med. 2005;352(4):380-91. [PubMed] - 4.
Mark BL, Vocadlo DJ, Oliver A. Providing beta-lactams a helping hand: targeting the AmpC beta-lactamase induction pathway. Future Microbiol. 2011;6(12):1415-27. [PubMed] - 5.
Shahid M, Malik A. Sheeba. Multidrug-resistant Pseudomonas aeruginosa strains harbouring R-plasmids and AmpC beta-lactamases isolated from hospitalised burn patients in a tertiary care hospital of North India. FEMS Microbiol Lett. 2003;228(2):181-6. [PubMed] - 6.
Juan C, Macia MD, Gutierrez O, Vidal C, Perez JL, Oliver A. Molecular mechanisms of beta-lactam resistance mediated by AmpC hyperproduction in Pseudomonas aeruginosa clinical strains. Antimicrob Agents Chemother. 2005;49(11):4733-8. [PubMed] - 7.
Sadikot RT, Blackwell TS, Christman JW, Prince AS. Pathogen-host interactions in Pseudomonas aeruginosa pneumonia. Am J Respir Crit Care Med. 2005;171(11):1209-23. [PubMed] - 8.
Puntarulo S. Iron, oxidative stress and human health. Mol Aspects Med. 2005;26(4-5):299-312. [PubMed] - 9.
Church D, Elsayed S, Reid O, Winston B, Lindsay R. Burn wound infections. Clin Microbiol Rev. 2006;19(2):403-34. [PubMed] - 10.
Paul M, Silbiger I, Grozinsky S, Soares-Weiser K, Leibovici L. Beta lactam antibiotic monotherapy versus beta lactam-aminoglycoside antibiotic combination therapy for sepsis. Cochrane Database Syst Rev. 2006;(1):CD003344. [PubMed] - 11.
Samaha-Kfoury JN, Araj GF. Recent developments in beta lactamases and extended spectrum beta lactamases. BMJ. 2003;327(7425):1209-13. [PubMed] - 12.
Infections due to Pseudomonas Species and related organisms. In: Kasper DL, editor(s). Harrisons Principles of Internal Medicine. 16th ed. New York: MCGrow Hill; 2005. - 13.
Dumas JL, van Delden C, Perron K, Kohler T. Analysis of antibiotic resistance gene expression in Pseudomonas aeruginosa by quantitative real-time-PCR. FEMS Microbiol Lett. 2006;254(2):217-25. [PubMed] - 14.
Fisher JF, Meroueh SO, Mobashery S. Bacterial resistance to beta-lactam antibiotics: compelling opportunism, compelling opportunity. Chem Rev. 2005;105(2):395-424. [PubMed] - 15.
Rice LB. The clinical consequences of antimicrobial resistance. Curr Opin Microbiol. 2009;12(5):476-81. [PubMed] - 16.
Ambler RP. The structure of beta-lactamases. Philos Trans R Soc Lond B Biol Sci. 1980;289(1036):321-31. [PubMed] - 17.
Jaurin B, Grundstrom T. ampC cephalosporinase of Escherichia coli K-12 has a different evolutionary origin from that of beta-lactamases of the penicillinase type. Proc Natl Acad Sci U S A. 1981;78(8):4897-901. [PubMed] - 18.
Jacoby GA. AmpC beta-lactamases. Clin Microbiol Rev. 2009;22(1):161-82. Table of Contents [PubMed] - 19.
Quale J, Bratu S, Gupta J, Landman D. Interplay of efflux system, ampC, and oprD expression in carbapenem resistance of Pseudomonas aeruginosa clinical isolates. Antimicrob Agents Chemother. 2006;50(5):1633-41. [PubMed] - 20.
Cabot G, Ocampo-Sosa AA, Tubau F, Macia MD, Rodriguez C, Moya B, et al. Overexpression of AmpC and efflux pumps in Pseudomonas aeruginosa isolates from bloodstream infections: prevalence and impact on resistance in a Spanish multicenter study. Antimicrob Agents Chemother. 2011;55(5):1906-11. [PubMed] - 21.
Xavier DE, Picao RC, Girardello R, Fehlberg LC, Gales AC. Efflux pumps expression and its association with porin down-regulation and beta-lactamase production among Pseudomonas aeruginosa causing bloodstream infections in Brazil. BMC Microbiol. 2010;10:217. [PubMed] - 22.
Tomas M, Doumith M, Warner M, Turton JF, Beceiro A, Bou G, et al. Efflux pumps, OprD porin, AmpC beta-lactamase, and multiresistance in Pseudomonas aeruginosa isolates from cystic fibrosis patients. Antimicrob Agents Chemother. 2010;54(5):2219-24. [PubMed]